An important consideration in the field of leukemia stem cells (LSCs) is to define culture condition that best mimic the in vivo microenvironment. LSCs rapidly differentiate when introduced in culture. This hampers our ability to conduct cell-based high-throughput lethality screens and partly explains the lack of success in drug development for this disease.
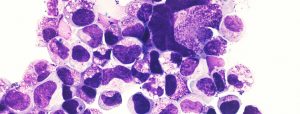
To overcome these constraints we conducted a high-throughput screening and identified small molecules that inhibit differentiation and support LSC activity in vitro. The first class of compounds identified, represented by SR1, inhibits the aryl hydrocarbon receptor (AhR) pathway. Interestingly this pathway is rapidly upregulated when cells are introduced in culture with obvious and rapid cell differentiation. A second compound, UM729, in combination with SR1 was shown to better maintain LSC activity in vitro. UM729 lacks AhR suppressor activity, which indicates that at least 2 different pathways are involved. These 2 compounds define the best culture conditions for improved ex vivo culture of primary human AML cells (Pabst et al., Nature Methods 2014).
Unfortunately, approximately 30% of specimens respond poorly to these molecules suggesting that a 3rd pathway is operational in securing LSC self-renewal divisions. That is why we continue to screen new molecules and investigate our best hits.
 The phenotypical differences arising from minor variations at the molecular level could have major effect on responses of leukemias to drugs. To address this issue we are screening the in vitro biological responses of primary human acute myeloid leukemia (AML) cells to various collections of chemical compounds in a media that transiently inhibits differentiation and supports leukemia stem cell activity ex vivo.
The phenotypical differences arising from minor variations at the molecular level could have major effect on responses of leukemias to drugs. To address this issue we are screening the in vitro biological responses of primary human acute myeloid leukemia (AML) cells to various collections of chemical compounds in a media that transiently inhibits differentiation and supports leukemia stem cell activity ex vivo.
Combination of biological responses to compounds with clinical, molecular and genetic data show that:
- cellular responses to some compounds correlate with clinical outcome
- some clinically approved drugs selectively inhibit only a fraction of specimens
- AML subgroups sharing a mutated pathway exhibit a distinct subgroup-specific compound responses
- interspecimen response variations identify compound clusters suggesting existence of a previously not identified shared pathway
In addition to improving prognostic stratification of AML our chemogenomic approach enables identification of novel compounds that selectively kill AML cell and identification of pathways that might represent novel therapeutic targets (Baccelli et al., Blood Cancer, 2017).
Antibody-based immunotherapies are particularly attractive as they produce antitumor activity without the burden of systemic toxicities encountered with conventional therapies.
encountered with conventional therapies.
Compared to solid tumors, antibody-based immunotherapy is underexploited in acute myeloid leukemia (AML), likely due to the limited availability of targetable surface antigens specifically expressed on AML cells.
Using a proteomic approach that we optimized for primary AML specimens, we are mapping the surface proteome (surfaceome) of AML subsets associated with poor prognosis to identify new AML-associated antigens.
This work is also accompanied by the development of companion tests for precision medicine, which will lead to the design of novel immunotherapies in AML.
 Our role is to develop or assist in the development of predictive tools that will make use of clinical observations and chemogenomic measures of patient samples to provide guidance to clinicians in selecting treatment. For this, we favor a two-pronged approach.
Our role is to develop or assist in the development of predictive tools that will make use of clinical observations and chemogenomic measures of patient samples to provide guidance to clinicians in selecting treatment. For this, we favor a two-pronged approach.
First, we emphasize the role of presentation tools to directly empower clinician-researchers to mine the data in the most transparent way possible.
In parallel, we apply modern machine learning techniques to derive clinically useful prediction models and thus contribute to the development of a more formal model of data integration.
As a foundation to the development of predictive tools, we have also been constantly refining an RNA-Seq analysis pipeline, providing information regarding gene expression levels and sequence variants (SNPs, indels and fusions) but also reporting on novel genes and quantifying splice variants.
 Current prognostic classification of Acute Myeloid Leukemia (AML) is based on cytogenetics and a limited number of mutations, where patients belong either to favorable, intermediate and adverse cytogenetic groups. The majority of patients will be classified into intermediate cytogenetics, an heterogenous group in which clinical outcome is highly variable and for which clinical decisions are difficult. This problem still leads to over- or under-therapy with dire consequences. In addition, a subset of patients with adverse cytogenetics will fail to respond to current therapies. More discriminatory risk stratification will lead to more accurate clinical decisions.
Current prognostic classification of Acute Myeloid Leukemia (AML) is based on cytogenetics and a limited number of mutations, where patients belong either to favorable, intermediate and adverse cytogenetic groups. The majority of patients will be classified into intermediate cytogenetics, an heterogenous group in which clinical outcome is highly variable and for which clinical decisions are difficult. This problem still leads to over- or under-therapy with dire consequences. In addition, a subset of patients with adverse cytogenetics will fail to respond to current therapies. More discriminatory risk stratification will lead to more accurate clinical decisions.
The Leucegene project aims at improving prognostic classification of AML using RNA-sequencing of a large cohort of BCLQ AML samples (n=457) from diverse cytogenetic subgroups. First, detailed mutational analysis is being performed to explore the landscape of known and novel mutations in AML. Transcriptomes are then correlated to clinical, cytogenetic and mutational data to identify the top differentially expressed genes to develop a transcriptomic-based classification.
As an exemple, EVI1-rearranged (EVI1-r) AML are characterized by distinct molecular alterations. We thus performed RNA sequencing of 12 EVI1-r AMLs and compared the results with those of other AML subtypes and normal CD34+ cells. EVI1-r AMLs have recurrent mutations in RAS and other signaling genes, splicing factors, and at a lower frequency, IKZF1 and TP53. EVI1-r AMLs also show a charactheristic transcriptome profile marked by high expression of MECOM, PREX2, MYCT1, PAWR and VIP (Lavallée VP et al. Blood 2015).
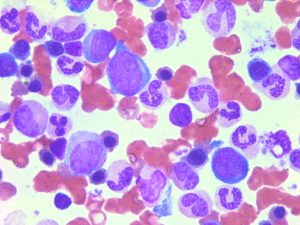 After induction or consolidation chemotherapy, patients in morphological complete remission can still have residual leukemic cells (i.e. minimal residual disease, MRD). MRD positivity is associated to an increased risk of relapse and a poorer clinical outcome. Currently, MRD is tested either by flow cytometry (leukemia- associated immunophenotype) or by RT-qPCR (aberrantly expressed gene, fusion transcript or mutation). However, a good MRD test is lacking in several AML subgroups.
After induction or consolidation chemotherapy, patients in morphological complete remission can still have residual leukemic cells (i.e. minimal residual disease, MRD). MRD positivity is associated to an increased risk of relapse and a poorer clinical outcome. Currently, MRD is tested either by flow cytometry (leukemia- associated immunophenotype) or by RT-qPCR (aberrantly expressed gene, fusion transcript or mutation). However, a good MRD test is lacking in several AML subgroups.
To address this issue, we are using RNA-sequencing data from our large cohort of AML samples and comparing it to sorted cell populations from normal peripheral blood and bone marrow specimens, to identify new leukemia-specific genes, which could represent better MRD markers.

Research resources and scientific data generated through the Leucegene project are made accessible to the scientific community after publication through international repositories.
The Leucegene RNA sequencing dataset (691 AML samples) is available in the Gene Expression Omnibus repository (GEO accession number GSE232130). Non-identifiable clinical data for the Leucegene cohort are available to academic investigators for a research project in hematological cancers approved by a research ethics committee in accordance with the procedures of the Quebec Leukemia Cell Bank: Banque de cellules leucémiques du Québec – Request for Cells/Data (bclq.org).
 RNA sequencing data is accessible through GEO or SRA.
RNA sequencing data is accessible through GEO or SRA.
More specifically, the following datasets are available:
- Leucegene: AML sequencing – Accession # GSE49642.
- Leucegene: AML sequencing (part 2) – Accession # GSE52656.
- Leucegene: AML sequencing (part 3) – Accession # GSE62190.
- Leucegene: AML sequencing (part 4) – Accession # GSE66917.
- Leucegene: AML sequencing (part 5) – Accession # GSE67039.
- Leucegene: AML sequencing (part 6) – Accession # GSE106272.
- Leucegene: AML sequencing– Accession # GSE67040 (SuperSerie containing the 6 SubSeries above).
- Leucegene: ALL sequencing – Accession # GSE49601.
- Transcriptome of Primitive Human Hematopoietic Cells: A New Resource to Find hHSC-Specific Genes – Accession: # GSE48846 and #GSE51984.
- Transcriptome analysis of G protein-coupled receptors in distinct genetic subgroups of AML – Accession: GSE98310 .
- Whole exome sequencing of human leukemia: SRA – Accession: PRJNA358716.
Software and databases are made available through web interfaces or code repositories:
The Leucegene Web Portal gives an overview of the Leucegene cohort and provides data exploration functionalities. While still in active development, it is accessible for the community and serves as an access point to obtain insight into the data available in the Leucegene Project. USE IT
EPCY
A command-line tool developed to rank best Predictive Gene (PG) to improve the detection of biomarker by evaluating gene expression on criteria aligned on the goal pursue, a.k.a their potential as predictive biomarkers. Publication SOURCE CODE DOC USE IT
km
Command-line tool developed to identify and quantify single nucleotide variants, insertions, deletions, and duplications from RNA-seq data (published in Life science Alliance). USE IT
MiSTIC
Integrative software package designed to provide an interface to view and interrogate large datasets including data derived from sequencing, clinical files, and chemical compound responses. It is also published in Nucleic Acid Research Journal. USE IT
BiDRA
A user-friendly web interface for the inference of dose-response curves and efficacy metrics (Max/Min, IC50, Slope) using Bayesian Inference through a Markov chain Monte Carlo approach (MCMC). The use of values distributions rather than single-point values allows for statistically sound analysis (Publication). USE IT
Chemical compounds generated through this program as prognostic tools or hits/leads for potential therapeutic applications are available once scientific data and patents are published.

 AML patient samples have been obained from the Quebec Leukemia Cell Bank. Clinical data is available through publications of each AML genetic subtype such as EVI1, MLL, Complex karyotype, Core Binding Factor rearrangements, CEBP mutations etc.
AML patient samples have been obained from the Quebec Leukemia Cell Bank. Clinical data is available through publications of each AML genetic subtype such as EVI1, MLL, Complex karyotype, Core Binding Factor rearrangements, CEBP mutations etc.
